The internet of things connects devices across the globe. Now researchers are considering how bacteria can join the network.
by Emerging Technology from the arXiv
Imagine designing the perfect device for the internet of things. What functions must it have? For a start,
by Emerging Technology from the arXiv

Imagine designing the perfect device for the internet of things. What functions must it have? For a start,
- it must be able to communicate, both with other devices and with its human overlords.
- It must be able to store and process information.
- And it must monitor its environment with a range of sensors.
- Finally, it will need some kind of built-in motor.
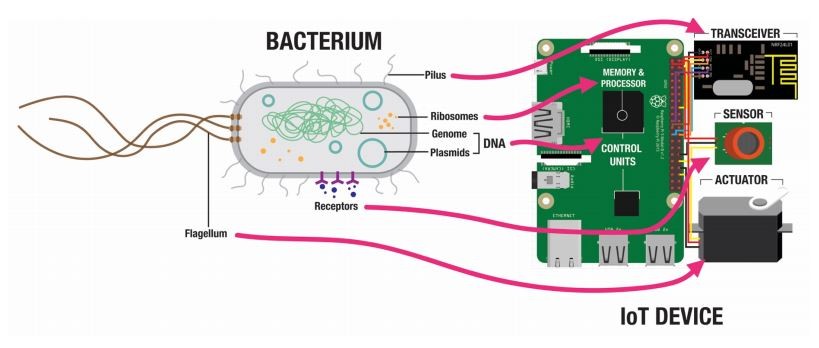
But another set of machines with similar functions is much more plentiful, say Raphael Kim and Stefan Poslad at Queen Mary University of London in the UK. They point out that bacteria communicate effectively and have built-in engines and sensors, as well as powerful information storage and processing architecture.
And that raises an interesting possibility, they say. Why not use bacteria to create a biological version of the internet of things? Today, in a call to action, they lay out some of the thinking and the technologies that could make this possible.
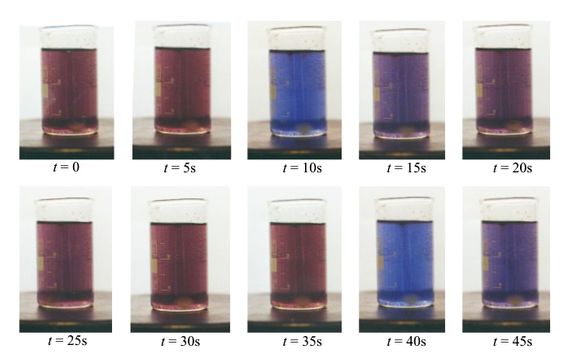
The way bacteria store and process information is an emerging area of research, much of it focused on the bacterial workhorse Escherichia coli. These (and other) bacteria store information in ring-shaped DNA structures called plasmids, which they transmit from one organism to the next in a process called conjugation.
Last year, Federico Tavella at the University of Padua in Italy and colleagues built a circuit in which one strain of immotile E. coli transmitted a simple “Hello world” message to a motile strain, which carried the information to another location.
This kind of information transmission occurs all the time in the bacterial world, creating a fantastically complex network. But Tavella and co’s proof-of-principle experiment shows how it can be exploited to create a kind of bio-internet, say Kim and Poslad.
E. coli make a perfect medium for this network. They are motile—they have a built-in engine in the form of waving, thread-like appendages called flagella, which generate thrust. They have receptors in their cell walls that sense aspects of their environment—temperature, light, chemicals, etc. They store information in DNA and process it using ribosomes. And they are tiny, allowing them to exist in environments that human-made technologies have trouble accessing.
E. coli are relatively easy to manipulate and engineer as well. The grassroots movement of DIY biology is making biotechnology tools cheaper and more easily available. The Amino Lab, for example, is a genetic engineering kit for schoolchildren, allowing them to reprogram E. coli to glow in the dark, among other things.
This kind of biohacking is becoming relatively common and shows the remarkable potential of a bio-internet of things. Kim and Poslad talk about a wide range of possibilities. “Bacteria could be programmed and deployed in different surroundings, such as the sea and ‘smart cities’, to sense for toxins and pollutants, gather data, and undertake bioremediation processes,” they say.
Bacteria could even be reprogrammed to treat diseases. “Harbouring DNA that encode useful hormones, for instance, the bacteria can swim to a chosen destination within the human body, [and] produce and release the hormones when triggered by the microbe’s internal sensor,” they suggest.
Of course, there are various downsides. While genetic engineering makes possible all kinds of amusing experiments, darker possibilities give biosecurity experts sleepless nights. It’s not hard to imagine bacteria acting as vectors for various nasty diseases, for example.
It’s also easy to lose bacteria. One thing they do not have is the equivalent of GPS. So tracking them is hard. Indeed, it can be almost impossible to track the information they transmit once it is released into the wild.
And therein lies one of the problems with a biological internet of things. The conventional internet is a way of starting with a message at one point in space and re-creating it at another point chosen by the sender. It allows humans, and increasingly devices, to communicate with each other across the planet.
Kim and Poslad’s bio-internet, on the other hand, offers a way of creating and releasing a message but little in the way of controlling where it ends up. The bionetwork created by bacterial conjugation is so mind-bogglingly vast that information can spread more or less anywhere. Biologists have observed the process of conjugation transferring genetic material from bacteria to yeast, to plants, and even to mammalian cells.
Evolution plays a role too. All living things are subject to its forces. No matter how benign a bacterium might seem, the process of evolution can wreak havoc via mutation and selection, with outcomes that are impossible to predict.
Then there is the problem of bad actors influencing this network. The conventional internet has attracted more than its fair share of individuals who release malware for nefarious purposes. The interest they might have in a biological internet of things is the stuff of nightmares.
Kim and Poslad acknowledge some of these issues, saying that creating a bacteria-based network presents fresh ethical issues. “Such challenges offer a rich area for discussion on the wider implication of bacteria driven Internet of Things systems,” they conclude with some understatement.
That’s a discussion worth having sooner rather than later.
Ref: arxiv.org/abs/1910.01974 : The Thing with E. coli: Highlighting Opportunities and Challenges of Integrating Bacteria in IoT and HCI.
ORIGINAL: MIT Technology Review
By Michael Schiffer / unsplash
Nov 1, 2019
Nov 1, 2019